Q&A with the scVerse team
AMA-style interview with Lukas Heumos, Giovanni Palla, Isaac Virshup, and Gregor Sturm from the scVerse team
This is reposted from a Q&A we did in 2022. Check out our Slack for more.
We had the chance to interview Lukas Heumos, Giovanni Palla, Isaac Virshup, and Gregor Sturm about the development of the popular single-cell python library scVerse. Lukas, Giovanni, Isaac, and Gregor are all on the @scverse team and below they discuss single-cell data analysis, open-source packages, upcoming scVerse functionalities, data visualization, and more!
Adam: I’ll get the scverse q+a started!
Question #1:
Adam: what are the most exciting frontiers for software development in single-cell omics? i.e., what emphasis is needed to help deal with the increasing scale of the data? anything interesting on the methodological side?
Giovanni: Imho i think a lot of potential is around libraries for model pre-training and deployment in useful way. i think huggingface is clearly leading in the non-bio space , in bio there is a lot of cool stuff (kipoi , bioimage.io ) but i feel there could be more for different applications and better ux (there is a nice thread on ux for bio right here: https://bitsinbio.slack.com/archives/c02rkfyrg3g/p1660672886575649 )
Anonymous: Sorry for newbie question but what would you be pretraining for exactly?
Giovanni: Great question, pre-training could be on large sc atlases to learn model able to perform zero-shot learning on various tasks. i think (1) which pretraining strategy to use, and (2) what to do with the pre-trained model are open questions. things that have been suggested in literature for (1) is reconstruction via autoencoders-type models and (2) is cell-type classification and query-to-reference dataset integration. i believe @adam has both ideas and ongoing projects on this.
Question #2:
Yohann: The diversity of single cell analysis and tools in the ecosystem, like cell ranger, starsolo or seurat, make it difficult to know where to start for new comers in the field. where does scverse fits in this landscape and how does it help new users?
Isaac: I'll answer this in two parts.
re: where does scverse fits in this landscape?
scverse focusses on libraries for the analysis of this data. broadly, i'd say this is everything downstream of the aligner. e.g. quite similar to seurat, except we're a broader community effort as opposed to a tool made by a single lab.
re: how does it help new users?
so many ways!
currently:
• we collect and encourage the development of learning resources. for example see our "learn" page, youtube channel , and the documentation of our projects.
• we provide a discourse forum for user questions
but we're working on plans for more. this includes organizing online workshops and tutorials.
you can read a bit more about our vision here from the "user engagement" section of our mission statement.
Lukas: I can add here that (not scverse project!) my lab is writing a book about single-cell analysis in all forms: https://github.com/theislab/cross-modal-single-cell-best-practices/
it makes heavy use of scverse tools due to the scalability and user friendliness of our tooling. i am confident that this will help people not only learn single-cell analysis, but as a side benefit also how to use scverse tooling.
Gregor: To bridge the gap to the preprocessing tools "upstream" of scverse, we are working together with the nf-core community to ramp up the scrnaseq workflow. the goal is to provide a universal workflow for the most common (pseudo-)aligners (kallisto/bustools, alevin-fry, starsolo and cellranger) that produces output in a format that can be readily imported by scverse tools.
Yohann: Thanks @gregor, we are actually deploying that specific nfcore workflow internally so i look forward to onboard the scverse!
Question #3:
Anonymous: As someone that is more bit than bio can you talk a bit about single cell analysis, how it helps researchers discover drugs that cant be done in other ways, and where scverse fits in
Gregor: I'll try to answer with an example from immuno-oncology, i.e. making drugs that leverage the body's immune system to fight cancer.
there are dozens of different immune cells types with different functions, and each of them may have different cell states. compared to "bulk" analyses, where a mixture of all cell-types is profiled together, with single-cell analysis you just get a higher resolution. this can be taken to the next level with the currently emerging "spatial" technologies, which profile cells together with their position in the tissue of origin. this is commonly illustrated using the smoothie-analogy.
to be a bit more concrete, single-cell analyses are useful in basically all stages of drug development, but particularly in early ones, for instance:
1. basic research. despite a lot of progress it is still poorly understood how most immune cells work together. recently, several atlases with 1m cells have been published that provide a comprehensive overview of immune cell-types and states in healthy and diseased tissue.
2. target selection: say you want to leverage a certain immune cell population for your drug. single-cell data can help to screen for potential target molecules that are expressed on the cell-type of interest.
3. indication prioritization: you already have a drug candidate that looks promising in pre-clinical experiments. using single-cell data you can select a patient cohort for a clinical trial that has a certain cell-type composition in which you expect the drug to work particularly well.
Anonymous: And the output is some read of transcription expression for each cell type?
Gregor: In the most common case, you get a matrix of cells x genes with the number of measured transcripts. typically, similar cells (i.e. cell-types or states) are grouped together and analyzed jointly.
i fell this review paper gives a nice overview of the different analysis steps if you are interested in more details:
https://www.nature.com/articles/s41592-021-01171-x
Question #4:
Nicholas: As people who work on prominent open source package in this space, how do you think about building open source software in bio? any advice for newcomers?
Isaac: a lot :rolling_on_the_floor_laughing:
i'll try and give a succinct answer here, but could definitely talk a lot about this!
it's getting better! the importance of open source is the space is getting more and more acknowledged, and funding directly for development seems to be getting more common.
it's also nice that bio software problems look more and more like problems from other fields. bio software has often felt closed off from the broader ecosytem (with entirely separate package repositories like `bioconda`, `bioconductor`). it's nice to rely on existing tools and ecosystems, and not having to build everything yourself!
career paths also seem clearer these days. whether with existing companies that appreciate open source (shoutout to genentech, boehringer), research software engineer positions, and open source startups.
Lukas: In my opinion working on os in bio is very rewarding. not only difficult technical challenges, but our daily work has the potential to contribute small pieces to find cures. i could not see myself working in unethical industries and am happy that there is this awesome space to do interesting and great work.
i also have the impression that academic labs are slowly starting to learn that good and well maintained software infinitely increases the visibility and the number of citations of a high impact paper. labs are more willing to hire rses and computationally talented phd students/postdocs to ensure exactly this.
Nicholas: If i were a software dev with little to no bio experience, where could i look to find an open source project to contribute to?
Isaac: This is a tough question. i've only really contributed to libraries that i was using. my best advice is to figure out what area of biology/ computer science you are interested in, and to go from there.
separately, there are many bio packages which don’t take pull requests. i think it’s just a bio authorship/ overworked comp bio thing. people who do take pull requests include:
scverse (ml for single cell, hi), mdanalysis (protein structure), nf-core (pipelines/ best practices), napari (python and qt based image viewer), open2c/ higlass (chromatin availablility analysis and web-based-viewers), and galaxy (no-code cloud platform for comp bio).
Gregor: The nf-core community (focused on implementing bioinformatics workflows in nextflow) is doing a great job in welcoming newcomers and organizing hackathons. both are things we would like to achieve with scverse, too, eventually.
other than that i would second what isaac said: start using libraries that look reasonably well-maintained, identify pain points, and get in touch with the developers on github.
Lukas: Especially for young academics (students, phd students) i want to emphasize that bio labs often times welcome them with open arms. i've already trained a couple of students that had 0 bio experience, but the right motivation. they quickly become strong programmers and learned the required bio on the side. i am only weary of recruiting pure tech people for data analysis projects since they require a much stronger biology background.
Isaac: Bio labs often times welcome them with open arms
this is really important. the best place to start is somewhere you can get a mentor.
Question #5:
Selin: Will scverse include methods/packages for integration of different datasets?
scverse seems to emphasize scale of the data, and i’m thinking of projects like the human lung cell atlas / https://www.biorxiv.org/content/10.1101/2022.03.10.483747v1|sikkema et al, biorxiv, 2022 showing that correctly applying integration methods - and evaluating the integrations - is important for being able to leverage the growing number of reference datasets being generated from different labs, projects, etc. wondering if it will be possible to integrate & evaluate the integrations of these datasets within scverse tools?
Isaac: Yes! the core package `scvi-tools` is widely used for this – and was used in the lung cell atlas project.
we do want to extend more in this area!
Lukas: The tool that we used to evaluate the integrations for lisa's hlca is scib, which provides a pipeline to evaluate several integration tools. the fundamental input datastructure is scverse's anndata.
generally, we are very happy to see people building scverse ecosystem tools that make use of our general purpose tooling. that's the whole idea.
Selin: Thanks so much isaac and lukas, great to hear and looking forward to scverse more!
Question #6:
Nicholas: How did each of you get involved in bio/open source/single cell/scverse?
Isaac: I did a bunch of stuff in undergrad, but settled on bio by my third year. i ended up doing my thesis on protein structure in a computational geometry lab where i learned python, and made my first pull request about six months later.
to me, open source is like an idealized version of science. people collaborating on cutting edge methods, having great open discussions, and making immediate visible impact on the field.
i eventually started a phd working on single cell transcriptomics. i became interested in scanpy and anndata because
• i saw the importance of having an interoperable data format
• i wanted to help bring python's great machine learning ecosystem to the field of single cell
• i just don't vibe with `r`
since i found open source so rewarding, and didn't really have an interest in the traditional academic career path, i spent more time working on these projects than on my phd (a tale as old as open source). now i'm on "hiatus" from the phd and am working full time as a software engineer on these projects!
Lukas: My career so far has been very straight forward with a bioinformatics bsc. and msc. very early on i joined a local bioinformatics core facility which put a heavy emphasis on building their own computational infrastructure (good decision!), which not only allowed me to contribute and learn a lot, but also exposed me to all kinds of software projects in bio.
eventually i learned about transcriptomics and machine learning and was looking for a way to combine those efforts. i joined the theislab and noticed that our computational tooling is growing so quickly in popularity and complexity that we needed a organization to manage that. luckily we had a couple of people around that have the same opinion and we joined forces to found scverse.
Giovanni: Started during phd and was just really lucky to have people guiding me at the beginning on how to do various kind of contributions. one great recent blogpost i found about how to get started with open source is this: https://merveenoyan.medium.com/open-source-contributio-how-to-get-started-9c149fde5a6
think it’s important to realize that contribution means answering issues, join discussions on various platform, submit documentation, and it’s a great way to start!
Adam: I went on a rather circuitous route that i don’t recommend, but also don’t regret. i went from operations research with internships in finance during college to cs with a ml focus to comp bio.
for open source we saw it as an opportunity to accelerate our phd research. we took great inspiration from scanpy which was a really important entry point for us to learn how to actually run an open source repo.
Question #7:
Brandon: How is scverse going to evolve as spatial omics become more population?
Giovanni: Lots of things going on in scverse for spatial omics data, we’ll hope to share prototypes soon (before end of year). the main project that is keeping us busy is the development of a new data format and in-memory representation that could support the *diversity* and *size* of spatial omics data.
after releasing squidpy, we realized that we couldn’t really handle all the complexity of spatial omics data in an efficient and intuitive way by just combining anndata and *a* image representation.
hence, we have been very lucky to team up with other developers from the imaging community (josh moore, leading ome-ngff, kevin yamauchi, core dev of napari, luca marconato, phd student embl) to build what we call `spatialdata`. we hope the project will enable spatial omics data storage and analysis by building open software and standards that join the imaging (napari, ome) and single-cell (scverse) communities.
for a high level overview, luca presented the project few months ago at the emblxjanelia seminar series, video
Question #8:
Brandon: How do you feel about the current technology behind single cell proteomics? are you excited about supporting it in the future?
Lukas: My lab (theislab) is currently working hard with mathias mann's lab to improve the experimental technology to make it work. our opinion is that real biological datasets are still extremely, extremely sparse and it is not the right time to develop scverse extensions for sc-proteomics yet. however, this can change very quickly and i am absolutely positive that someone in our lab will pick this up.
so, yes!
Question #9:
Nicholas: How does the tooling for single cell workflows differ from bulk rna-seq workflows? are there common abstractions that you can use?
Isaac:
bigger datasets
since observations are now cells, instead of samples, you have a lot more of them. broadly this means you need to worry more about data structures and performance. for instance, not storing your 1 million cell by 20 thousand gene matrix in a csv.
more unknowns
bulk studies were very well structured (or at least, the useful ones were). you basically knew everything about the samples up front, and were interested in what the features were doing in that design.
there's a lot less we know about each observation in most single cell datasets (at least droplet based ones). while we do know what sample a cell came from, we don't know what cell type it is. this makes tools for clustering and classifying cell types much more important.
a cell is a fundamental unit, a sample is not
not only do have to do more work around the unit of observation, it's a more useful unit of observation. it's very difficult to draw equivalencies between samples from different bulk studies. were samples collected in similar ways? were similar flow panels used?
with single cell, we can broadly say a cell in person a should have an equivalent in person b. this means we can actually do integrative analysis.
Lukas: I would like to add that it is possible to leverage the single-cell information for bulk experiments by deconvolving cell types for bulk data. there's a wide range of tools available for this purpose.
Question #10:
Nicholas: How do you think about visualizing single cell data? this preprint was about a year ago:

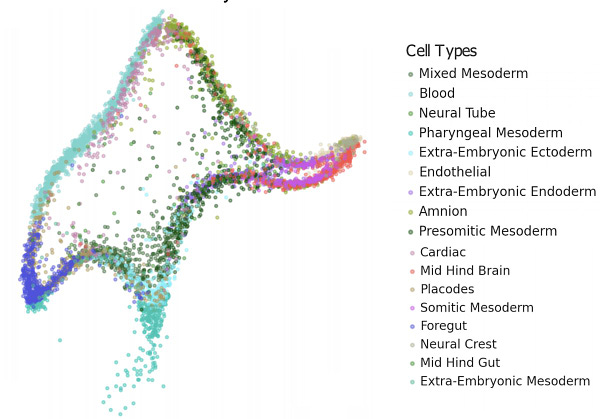
— did it affect how you look at your data?
Jake: My rule of my thumb, if the data has structure it should be immediately obvious. pca then umap is a reasonable place to start. never a good idea to fiddle parameters until you find what you're looking for.
Isaac: I think that the paper makes some good points about umap not giving you meaningful dimensions. and as the field has grown i've definitley seen some... questionable interpretations of umap/ tsne embeddings.
however, graph based methods have worked well for single cell data. they've proved effective for clustering, integration, and fate mapping. since we use this graph representation for so many parts of the analysis, it's important to be able to inspect it.
Adam: I agree with isaac, also as the scale of the data grows, these types of visualizations become even less useful in my opinion. i’m sure in the future we will have a suite of standard metrics and analyses that help replace using umap as a basis for hypothesis generation.
Nicholas: What are the standard visualizations that you use? is this part of scverse?
Adam: Well, i’ve used a lot of umaps in my day :upside_down_face:, but scanpy has really convenient plotting tools for heatmaps, dotplots, violin plots, and much more. https://scanpy-tutorials.readthedocs.io/en/latest/plotting/core.html
Isaac: cellxgene has been invaluable for my work. it's very well made interactive viewer for single cell datasets. you can just point it at your anndata object! it makes vizualiting the data available to a non-technical audience.
vitessce is another cool tool in this area, and is entirely browser based!
Travis: Sorry i am late to the party. i would add to @jake statements “if the data has structure it should be immediately obvious”. yes, but … only if you have a systematic and justifiable means of processing the raw data. did you filter your data to remove spurious samples? how did you handle zeros? did you do any transformations to the data like standardization. with single cell data it is common to log transform and the decision of log(x+1) vs log(x+\epsilon) can change some analysis. i think what makes comp-bio so hard is that you have to be proficient and understand how the data was collected and be a “good” data scientist, know and understand ml and inference… and have you fingers on biology. the patterns don’t usually jump out at you if all of the steps in your pipeline aren’t carefully designed and justified
Question #11:
Akshay: Do/how do you think about extending the scvi-tools-style factor model toolkit to other modalities of biochemical data?
Adam: Great question! we aren’t actively pursuing this in the lab, but others have extended these models to micro biome data https://www.biorxiv.org/content/10.1101/2021.11.09.467939v1. what sorts of modalities did you have in mind?
Question #12:
Nicholas: What other open source packages do you all respect and like a lot?
Adam: As we develop ml models as part of scvi-tools, we have grown to really appreciate pytorch of course, but also pytorch lightning!! it’s been really helpful to cut tech debt in the form of old and less useful training loops. more recently we found that jax improved the speed of scvi significantly, so we are trying to become more familiar with that ecosystem.
Giovanni: One project that has grown tremendously in the last two years is napari and imho has been transformative for image analysis in python
Lukas: Nf-core inspired me a lot when it comes to community building, brand building and tooling harmonization.
Isaac: `bioconductor` is great. it's really pushed open source in the high througput sequencing field, and come up with a great model for it. high quality core tools which methods developers build on top of is a great idea. also their community is great!
`nf-core` has also done a really fantastic job with community building. and has accomplished the impossible: get people to agree on a workflow language!
`julia` is fantastic too. it's abstractions work very well for math-y stuff, but it also lets you get very low level – without having to use a statically compiled language.
one common thread for all these, they are explicitly collaborative. they're all a platform for you to build your own thing on, not a monolithic tool.
Nicholas: R or python :eyes:
Adam: …
Adam: Short answer: both are useful in their own ways, ggplot is way better than matplotlib, r has cool stats packages, but i think python has a more promising future in the single cell space as the data sizes massively increase.
Lukas: Actually, i will answer this with stats. in the single-cell community python is set to overtake r very soon and growing at a much more rapid pace.
Lukas: Source: zappia 2021
Giovanni: Thanks for bringing this up :sweat_smile: . imho a fair statement is r is better for visualizing tabular data (good luck with anything else that is not a table) and for doing any type of linear modelling (glm/glmm/…). anything else, from my experience, python has a richer/more performant alternative.
Isaac: Personally, i quite like julia.
buuut, i work in python because it's easy for people to learn, the unmatched machine learning ecosystem, and because it's "the second best language for everything". the later two are too strong of network effects to ignore!
Question #13:
Nicholas: How is scverse organized? is there a steering committee?
Lukas: We have several layers of organization. see: https://scverse.org/people/
every major package that we support gets at least one core developer as representatives that become part of our core team. the core team meets regularly and makes democratic decisions. major decisions or tie breaks are decided by the steering council which is always supposed to be an uneven number.
we also get support and input from major pis in the space that support us financially/with logistics (management committee) and advice (advisory committee).


